Technology focus: 10x Genomics Visium#
This notebook will present a rough overview of the plotting functionalities that spatialdata implements for Visium data.
Loading the data#
Please download the data from here: Visium dataset and rename it (eventually using symlinks) to visium_brain.zarr.
visium_zarr_path = "./visium_brain.zarr"
import spatialdata as sd
visium_sdata = sd.read_zarr(visium_zarr_path)
visium_sdata
SpatialData object with:
├── Images
│ ├── 'ST8059048_hires_image': SpatialImage[cyx] (3, 2000, 1969)
│ ├── 'ST8059048_lowres_image': SpatialImage[cyx] (3, 600, 591)
│ ├── 'ST8059050_hires_image': SpatialImage[cyx] (3, 2000, 1968)
│ └── 'ST8059050_lowres_image': SpatialImage[cyx] (3, 600, 590)
├── Shapes
│ ├── 'ST8059048': GeoDataFrame shape: (4992, 2) (2D shapes)
│ └── 'ST8059050': GeoDataFrame shape: (3497, 2) (2D shapes)
└── Tables
└── 'table': AnnData (8489, 31053)
with coordinate systems:
▸ 'ST8059048', with elements:
ST8059048_hires_image (Images), ST8059048 (Shapes)
▸ 'ST8059050', with elements:
ST8059050_hires_image (Images), ST8059050 (Shapes)
Visualise the data#
We’re going to create a naiive visualisation of the data, overlaying the Visium spots and the tissue images. For this, we need to load the spatialdata_plot library which extends the sd.SpatialData object with the .pl module.
import spatialdata_plot
visium_sdata.pl.render_images().pl.render_shapes().pl.show("ST8059050")
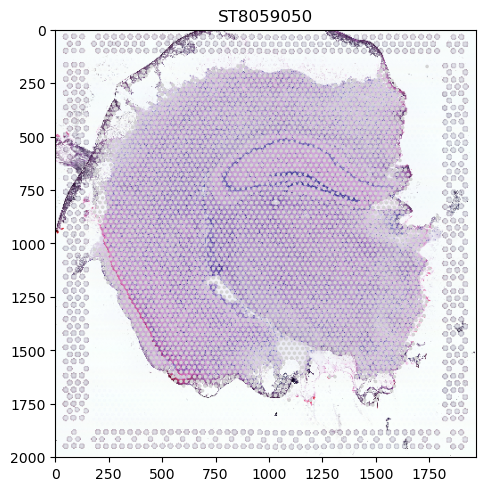
We can see that the data contains two coordinate systems (ST8059050 and ST8059052) with image and spot information each. In SpatialData, these spots are represented as Shapes. When giving no further parameters, one panel is generated per coordinate system with the members that have been specified in the function call. We can see that the spots are aligned to the tissue representation which is also respected by the plotting logic.
However, the spots are all grey since we have not provided any information on what they should encode. Such information can be found in the Table attribute (which is an anndata.AnnData table) of the SpatialData object, either in the data itself or the obs attribute.
visium_sdata["table"].to_df().sum(axis=0).sort_values(ascending=False).head(10)
# We will select some of the highly expressed genes for this example
mt-Co3 3366555.0
mt-Co1 3134006.0
mt-Atp6 2175621.0
mt-Co2 2160662.0
mt-Cytb 1317073.0
mt-Nd4 1100399.0
mt-Nd1 1098630.0
Fth1 840543.0
Ttr 833197.0
mt-Nd2 775515.0
dtype: float32
visium_sdata["table"].obs.head(3)
| in_tissue | array_row | array_col | spot_id | region | |
|---|---|---|---|---|---|
| AAACAACGAATAGTTC-1 | 0 | 0 | 16 | 0 | ST8059048 |
| AAACAAGTATCTCCCA-1 | 1 | 50 | 102 | 1 | ST8059048 |
| AAACAATCTACTAGCA-1 | 0 | 3 | 43 | 2 | ST8059048 |
Color the visium spots by gene expression#
To use this information in our plot, we pass the name of the column by which we want to color our expression to color. Furthermore, we are going to subset the data to only one coordinate system.
(
visium_sdata.pp.get_elements("ST8059050")
.pl.render_images(elements="ST8059050_hires_image")
.pl.render_shapes(elements="ST8059050", color="mt-Co3")
.pl.show()
)

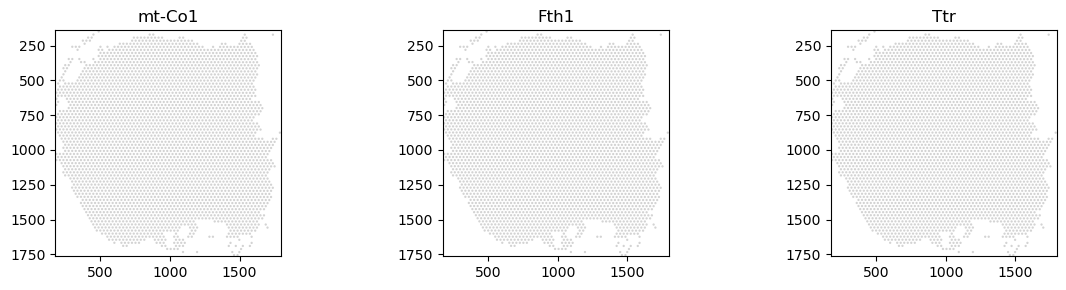
We can also provide ax objects to spatialdata_plot for further customisation.
import matplotlib.pyplot as plt
fig, axs = plt.subplots(ncols=3, nrows=1, figsize=(12, 3))
visium_sdata_subset = visium_sdata.pp.get_elements("ST8059050")
visium_sdata_subset.pl.render_shapes(elements="ST8059050", color="mt-Co1").pl.show(ax=axs[0], title="mt-Co1")
visium_sdata_subset.pl.render_shapes(elements="ST8059050", color="Fth1").pl.show(ax=axs[1], title="Fth1")
visium_sdata_subset.pl.render_shapes(elements="ST8059050", color="Ttr").pl.show(ax=axs[2], title="Ttr")
plt.tight_layout()